What is Multiplex qPCR in Cannabis Testing
The word “multiplex” changes everything about what a test
result tells you. A standard PCR asks one question. A
multiplex qPCR panel for three pathogens answers six
questions at once — each with a measurement, not just a yes or no.
What is Multiplex qPCR?
Multiple pathogens. One sample. One run. Quantitative load per target.
The only testing method that provides actionable diagnostic information before symptoms appear — with pathogen concentration data that drives real operational decisions.
How qPCR Works
Polymerase Chain Reaction (PCR) detects genetic material by amplifying it until it becomes measurable. If the target sequence is present in the sample — even at very low concentrations — PCR amplifies it exponentially and generates a detectable signal. If the target sequence is absent, no amplification occurs.
The amplification cycle
Denaturation
The DNA or RNA target sequence is separated by heat, unwinding the double strand and making it accessible for binding.
Primer binding
Short sequences (primers) designed to flank the target bind to each end of the region of interest.
Extension — billions of copies
DNA polymerase copies the target sequence. After 30–40 cycles, a handful of molecules becomes billions, producing a measurable fluorescent signal.
Real-time signal detection (qPCR)
Fluorescent probes measure amplification at each cycle — allowing not just detection, but measurement of how much pathogen is present.
RT-qPCR for RNA pathogens
For RNA-based pathogens — HLVd and Cannabis Cryptic Virus (CCV) — an extra step is required. Their RNA must first be converted into complementary DNA (cDNA) using reverse transcriptase. This is called RT-qPCR. DNA-based pathogens like fungi are detected directly with standard qPCR.
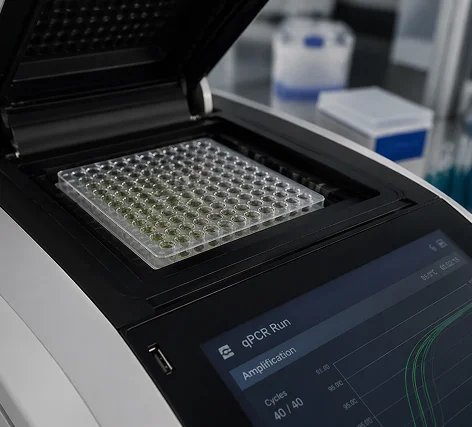

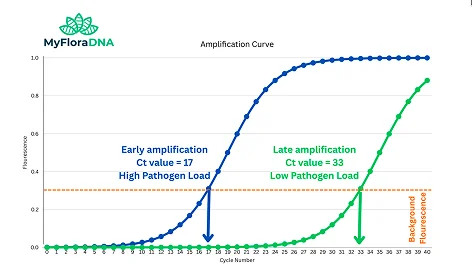
The Ct Value — Not Just Positive or Negative
In standard PCR, the result is binary. In quantitative PCR (qPCR), fluorescent probes produce a signal at each cycle, and the instrument measures how many cycles it took for that signal to cross a threshold. This is the Cycle Threshold (Ct value).
A low Ct value = target detected quickly = high pathogen load. A high Ct value = target detected late = low pathogen load. This single number is the difference between a result that says “positive” and one that tells you how serious the infection is.
Ct value is not just a technical detail. A Ct of 17 in a mother
plant means heavy, established infection — quarantine immediately. A Ct of 35 may represent early-stage infection
or low-level background — flag for retest. Binary results cannot give you this. The Ct value is what drives the triage decision.
Ct value reference guide
Ct <25 High Load
Heavy, established infection
Pathogen highly concentrated throughout plant tissue. Detected in early cycles.
→ Immediate quarantine + removal. Trace all cuttings taken from this plant.
Ct 25-32 Mod. load
Moderate pathogen load
Established infection confirmed. Do not take cuttings from this plant.
→ Remove from propagation immediately. Quarantine and re-evaluate.
Ct >32 Low load
Low pathogen load
May represent early-stage infection, residual signal, or sample-level variation.
→ Flag for re-test in 2–4 weeks. Do not propagate until status confirmed.
N/D Not detected
No amplification observed
Target absent or below limit of detection in the sample tested.
→ Re-test at next scheduled interval per your testing SOP.
Multiple targets, one run
Multiplexing means running multiple detection targets in the same reaction tube simultaneously. Each target is detected using different colored fluorescent probes so the instrument reads each signal independently and reports a Ct value for each pathogen in
the same run.
Pathogens detected by MyFloraDNA panels
Viroid
Hop Latent Viroid (HLVd)
Primary viroid affecting commercial cannabis. Asymptomatic for months.
Virus
Cannabis Cryptic Virus (CCV)
Common HLVd co-infector. Independently suppresses secondary metabolites.
Fungal
Fusarium oxysporum
Root pathogen. Early symptoms mimic environmental stress.
Fungal
Fusarium solani
Soil-borne fungal pathogen with similar presentation.
Water Mold
Pythium myriotylum
Root rot. Common in hydroponic systems. Silent until collapse.
Fungal
Botrytis cinerea
Gray mold. Primary post-vegetative and harvest risk.
A negative HLVd result doesn’t rule out CCV or Fusarium
In a facility where CCV is present alongside HLVd, a grower who only tests for HLVd may quarantine HLVd-positive plants while re-integrating CCV-positive plants that cleared the HLVd screen — and the quality problem continues.
Available panels
HLV Shield™
Proactive 3-in-1 screening for HLVd, Fusarium oxysporum, and Pythium myriotylum — the baseline test for any clean stock program.
Wilting Panel™
For plants that droop or collapse despite proper watering — rules out the three root and vascular pathogens most likely behind it.
Yellowing Panel™
For chlorosis and vein clearing that nutrients don’t correct — screens for the viral causes growers most commonly misattribute to deficiency.
Mosaic Panel™
For leaves showing mottled, patterned, or light/dark green discoloration — identifies the viral co-infections behind it in one run.
Leaf Deformation Panel™
For new growth that comes out twisted, curled, or structurally abnormal — screens the pathogen causes before blaming genetics or environment.
Leaf Spot Panel™
For brown or yellow spots, expanding lesions, and localized necrosis — confirms whether the cause is fungal before adjusting your protocol.
Flower Failure Panel™
For flowers that brown, collapse, or rot before harvest — identifies the fungal pathogens behind bud loss while there’s still time to act.
General Decline Panel™
For crops that grow but underperform with no clear cause — screens HLVd and the two Fusarium species most associated with silent productivity loss.
Single-Pathogen PCR vs. Multiplex qPCR
Compared across the dimensions that matter for commercial cannabis operations.
| Feature | Single-Pathogen PCR | Multiplex qPCR (MyFloraDNA) |
|---|---|---|
| Pathogens per run | 1 | 3 simultaneously |
| Result type | ✗ Positive / Negative only | ✓ Quantitative Ct load per target |
| Co-infection visibility | ✗ None | ✓ All targets quantified in one result |
| Cost per target | Higher — one assay per pathogen | ✓ Lower — shared run cost |
| Turnaround | Sequential — longer total wait | ✓ Single run — faster answers |
| Validated controls | Varies by provider | ✓ Positive + negative on every run |
| Triage information | ✗ Presence / absence only | ✓ Viral load guides quarantine decisions |
The cost-per-target advantage is significant. Running three separate single-pathogen PCR assays costs more in total than running one multiplex panel covering all three targets — and takes longer because sequential assays require sequential turnaround times. The multiplex approach delivers more information faster at lower per-target cost.
What Validation Means and Why It Matters
Not all qPCR tests are equivalent. A test is only as reliable as the validation work behind it. For a cannabis pathogen panel, validation means establishing four things.
Sensitivity — Limit of Detection (LOD)
The lowest pathogen concentration at which the assay reliably produces a positive result. Missing a low-load infection in a mother plant allows widespread facility contamination — the plant tests negative, stays in the room, and keeps distributing infected cuttings.
Specificity — No false positives
Whether the assay only amplifies the intended target. Confirmed by testing against a panel of related organisms and verifying only the intended target produces amplification.
Repeatability — Consistent across runs
Results must be reproducible across independent runs. A result that varies significantly between identical samples is not a reliable diagnostic tool — it is noise.
False negative risk — Sampling strategy
False negatives are a real risk due to uneven pathogen distribution within a plant. Samples should be collected from multiple plants per batch and multiple tissue types per plant. A negative result from a single sample does not guarantee absence of the pathogen.
Every MyFloraDNA run includes both controls
Run controls
A sample with a known quantity of the target pathogen. Confirms the assay is amplifying the target as expected.
Negative control
A sample with no target present. Confirms no contamination occurred during sample processing.
A negative result without a passing positive control is not a reliable negative. If either control fails, the run is invalid and results are not reported. Ask any testing provider whether they run controls on every plate.

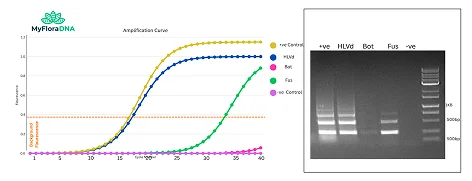
Amplification curves show clear detection of each target. Gel electrophoresis confirms target-specific amplification at expected product sizes. Positive control shows all targets; negative control shows no amplification.
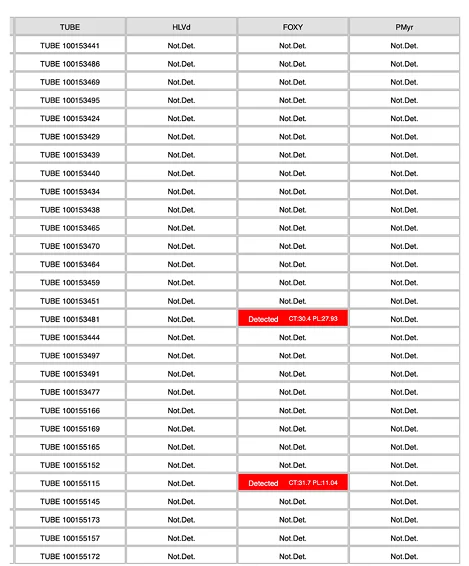
How to Read a MyFloraDNA Result
A MyFloraDNA multiplex panel result reports each target independently with a detection status and Ct value. Each result maps directly to an operational action.
Ct < 25
Heavy, well-established infection. Pathogen highly concentrated throughout tissue.
→ Immediate quarantine and removal. Trace all cuttings already taken from this plant.
Ct 25-32
Established infection confirmed. Plant is actively infected and capable of spreading through cuttings.
→ Remove from propagation immediately. Do not take cuttings. Quarantine and re-evaluate.
Ct > 32
May represent early-stage infection, residual signal, or sample-level variation.
→ Flag for re-test within 2–4 weeks. Do not propagate until status confirmed.
Not detected
Target is absent or below the limit of detection in the sample tested.
→ Re-test at next scheduled interval per your testing SOP.
Operational Context
For how to structure testing around these results — testing
cadence, quarantine protocols, tissue culture recovery — see the Cannabis Mother Room Biosecurity Guide.
Get quantitative pathogen data from one sample, one run.
Submit a sample and get results via MyFloraCLOUD in 72 hours or less.
Frequently Asked Questions
A validated assay with confirmed specificity produces very few false positives. The most common source is sample contamination during processing — which is why negative controls on every run are non-negotiable. If a negative control tests positive, the run is invalid. MyFloraDNA does not report results for invalid runs.
Typically a small amount of fresh leaf tissue is sufficient. MyFloraDNA provides sample submission instructions with each order. Tissue should be collected from actively growing material and stored appropriately for shipment to preserve RNA integrity.
Results are typically delivered in under 72 hours. Turnaround time may vary slightly based on sample volume and laboratory throughput. Expedited processing may be available for time-sensitive decisions.
MyFloraDNA’s HLVd assay targets conserved regions of the viroid genome, designed to detect the range of HLVd sequence variants documented in commercial cannabis. Primer and probe design is reviewed against published sequence data from cannabis HLVd isolates.
Regulatory acceptance of specific test methods varies by jurisdiction and program. Contact MyFloraDNA to discuss whether specific panels meet the requirements of your state or provincial clean stock certification program.
Resources
Related Guides
Diagnostics, biosecurity, and remediation guidance
